Research
Epithelial to Mesenchymal Transition (EMT)
Metastasis is a critical event in cancer progression that involves the spread of tumor cells from a primary cancer to secondary sites in the body, thus rendering the effective management of cancer extremely difficult. The misregulation of several normal biological pathways contributes to metastasis.
In particular, a process known as 'Epithelial to Mesenchymal Transition' (EMT), which causes cells to change their shape and migrate during development, is hypothesized to play an important role in metastasis. Our research aims to understand the epigenetic and molecular basis of EMT. Specifically, our lab is interested in two highly related transcription factors, Snail and Slug, which regulate the expression of genes that are essential for EMT.
The Snail and Slug transcription factors in EMT
Snail and Slug are primarily repressors of transcription that recognize and bind to the same minimal DNA sequence, CAGGTG, and elicit changes in the epigenetic landscape of genes by recruitment of co-factors such as the histone lysine demethylase LSD1, the polycomb repressive complex PRC2, and Histone deacetylases (HDACs). This causes cells to become migratory, invasive, and de-differentiated, leading to metastasis. In the image below, loss of E-cadherin (red), a cell-cell adhesion protein, following induction of EMT is seen.
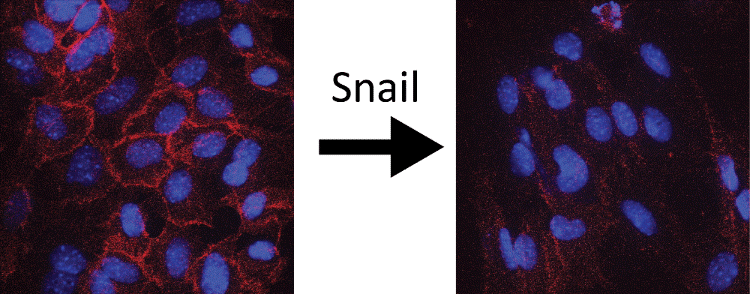
Using breast cancer cells as our model system, we propose to investigate the mechanisms utilized by Snail and Slug to bring about gene repression leading to EMT. As our understanding of epigenetic mechanisms of EMT increases, the challenge will be to develop new drugs to specifically target and reverse each epigenetic lesion.
The rationale for our proposed research is that detailed elucidation of the epigenetic mechanisms in EMT will provide valuable insights into the pathology of cancer metastasis, provide the basis for translational cancer research, and in the long term, will identify new targets for the personalized management of cancer metastasis.
Gene Regulation During EMT
The Epithelial to Mesenchymal Transition involves changes in cellular phenotype from a well-differentiated, immobile state to a non-adhesive, mobile phenotype. This transition is involved in early development, when cells migrate to form different germ layers and hence tissues, to cancer metastasis, when cells move from the original tumor site to other locations in the body. Our lab aims to understand the molecular basis for this transition, specifically addressing 3 important questions:
- What are the signals that induce EMT?
- How are EMT-related transcription factors such as SNAIL-regulated?
- What roles do transcription factor proteins play during EMT?
- Does each intermediate E/M state have a ‘memory’ of transcription?
Transcriptional priming in Lyme Arthritis
Lyme disease is a multisystem infection by the spirochetal bacterium Borrelia burgdorferi (Bb) that is spread by ticks. While often managed with antibiotics and anti-inflammatory drugs, a significant percentage of patients exhibit antibiotic-refractory Lyme disease, which persists for months to years after treatment in the absence of any signs of active infection, also referred to as post-treatment Lyme disease syndrome (PTLDS). It remains unclear why symptoms persist so many years after the initial infection, and there is an urgent need for new approaches to pinpoint underlying causes of PTLDS, for which few treatment options exist. We propose the novel idea that long-term propagation of epigenetic changes is the underlying cause of persistent symptoms, even after the original infection has cleared. As epigenetic changes are pharmacologically reversible, this would allow for new ways to treat PTLDS. We are collaborating on this endeavor with the Brissette Lab.
We are addressing the following questions:
- What are the epigenetic changes that occur in arthritic cells following infection with the spirochete Borrelia burgdorferi?
- What are the mechanisms that maintain the epigenetically primed, pro-inflammatory state following infection?
- Can we reverse epigenetic priming and reverse inflammation?
3D models of EMT and arthritis
While 2D cell culture models have been used as model systems for many decades, 3D model systems have emerged in recent years as better mimics of tissue and organ systems. We are collaborating with Dr. Ali Alshami’s group to develop novel biomaterials for EMT and arthritis research. These materials are designed for growing different cell types and will be tuned in various ways to optimize cell adhesion, stiffness, etc. for evaluating cellular responses to varied stimuli. This will allow us to model cell behaviors such as in vitro cell growth and differentiation, cell response to infection, cancer growth and metastatic progression, etc.