Adenosine Quantification
The standard curve
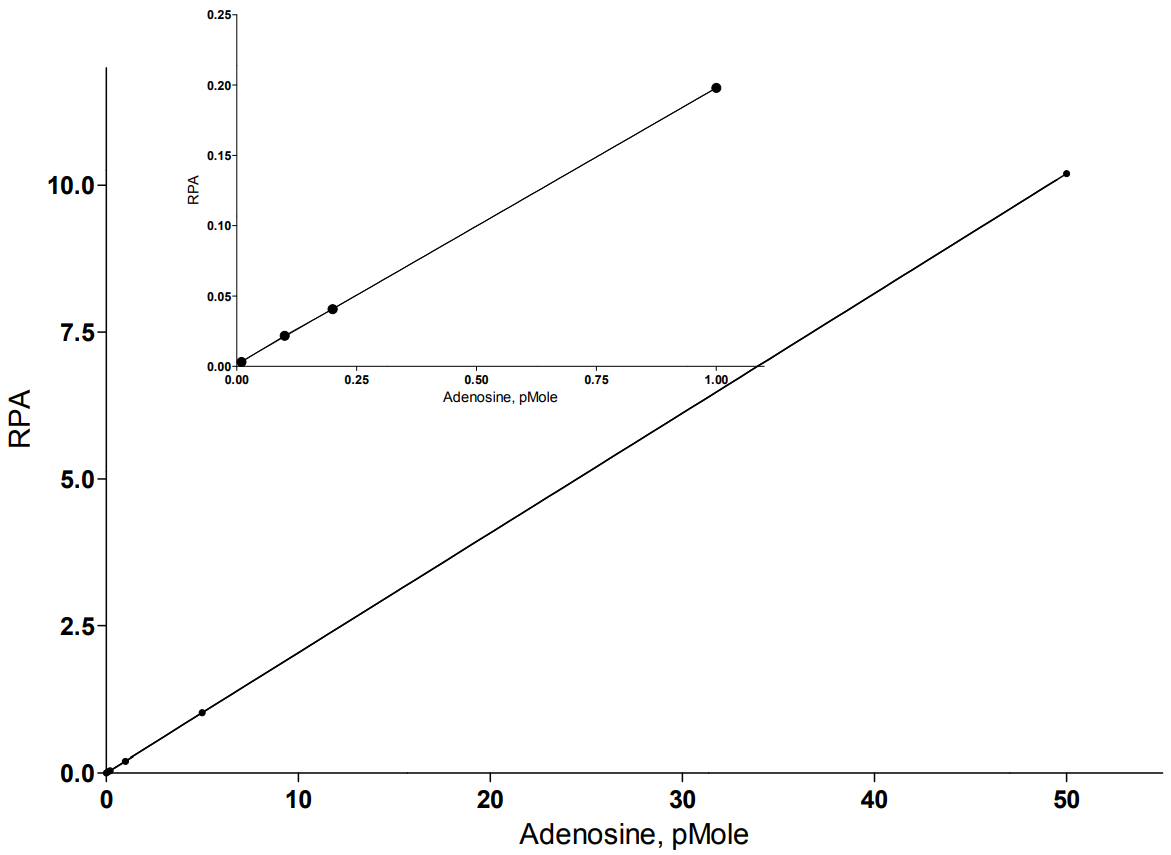
Reverse-phase LC electrospray ionization mass spectrometry method was developed for adenosine quantification. The HPLC was carried out using a Luna C-18(2) (3 μm column, 100 A pore diameter, 150 x 2.0 mm) (Phenomenex, Torrance, CA, USA) with security guard cartridge system (C-18) (Phenomenex, Torrance, CA, USA). The system was consisted of an Agilent 1100 series LC pump with wellplate autosampler (Agilent Technologies, Santa Clara, CA). The autosampler was set at 4o C, and a chromatographic column temperature was maintained at 35oC. A 10 μL of sample was injected onto a chromatographic column.
The solvent program for elution was the modification of a previous method (1-2). This modification was made to decrease separation time and increase sensitivity of detection by increasing peak sharpness. The LC system was composed of 25mM ammonium acetate in water (solvent A) and acetonitrile (solvent B). The separation was carried out at flow rate 0.2 ml/min and isocratic concentration of solvent B (10%). The total separation time was 5 min, and no equilibration between runs was required.
MS analysis was performed using a quadrapole mass spectrometer (API3000, Applied Biosystem,
Foster City, CA, USA) equipped with TurboIonSpray ionization source. Analyst software
version 1.4.2 (Applied Biosystem) was used for instrument control, data acquisition,
and data analysis. The mass spectrometer was optimized in the multiple reaction-monitoring
mode. The source was operated in negative ion electrospray mode at 450oC, electrospray voltage was 5000 V, nebulizer and curtain gas were 8 L/min. Declustering,
focusing, entrance,
focusing potential, and collision energy potentials were 30, 250, 10, and 30 V, respectively.
The quadrupole mass spectrometer was operated at unit resolution. Adenosine was quantified
against internal standard ribose-13C5-adenine (5pMole) added to brain tissue extract prior to analysis. Precursor/product
ions used for quantification were 267.8/136.2 for adenosine, and 272.8/136.2 for ribose-13C5-adenine.